Exact nearest-neighbor search is essentially impossible at scale. A billion 1024-dimensional vectors means a trillion floating-point operations per query, and you need an answer in milliseconds. Every production vector database makes the same trade-off: give up the guarantee of finding the exact match in exchange for finding an excellent match, fast.
Databases like OpenSearch, Pinecone, and QDrant handle this using specialized algorithms like IVF and HNSW. This post builds both from scratch to understand the accuracy/speed trade-off. It does not cover product quantization, ScaNN, or DiskANN. Familiarity with vector embeddings is assumed.
Measuring closeness
Given a search query, vector search finds the documents whose embeddings are closest to the query’s embedding. “Closest” requires a distance function. Two are standard:
- Euclidean distance measures how far apart two vectors are in space:
- Dot product measures the angle between two vectors scaled by their magnitudes:


Euclidean distance is the default in most vector databases and the metric used throughout this post. Dot product (or its normalized form, cosine similarity) is preferred when vector magnitudes are meaningful, such as in recommendation systems where a larger magnitude signals stronger preference.
Given a corpus of vectors and a query vector , k-nearest-neighbor (kNN) search finds the vectors closest to . The rest of this post is about making that selection fast.
Naive approach: The Linear Scan
The simplest kNN implementation is a linear scan: compute the distance from the query to every vector in the corpus, then return the smallest.
We call this a Flat Index.
class FlatIndex:
def __init__(self):
self._vectors = []
def index(self, vector: np.ndarray):
self._vectors.append(vector)
def search(self, query: np.ndarray, k: int) -> list[tuple[int, float]]:
# 1. Compute euclidean distance from query to every vector in the corpus
dataset_matrix = np.stack(self._vectors)
distances = np.linalg.norm(dataset_matrix - query, axis=1)
# 2. Partial sort: O(n) to get top-k, then sort only those k elements
nearest_indices = np.argpartition(distances, k)[:k]
nearest_indices = nearest_indices[np.argsort(distances[nearest_indices])]
# 3. Return (id, distance) pairs
return [(int(i), float(distances[i])) for i in nearest_indices]This works for small datasets. The problem is that it’s a brute-force scan.
If we have vectors of dimension , every search takes time. At 1 million vectors and 128 dimensions, that is 128 million multiplications per query. Production indexes routinely hold a billion vectors at 1024 dimensions.
Running a simple benchmark on a consumer laptop shows almost 1.2s per query at 1 million vectors and 128 dimensions.
> uv run plot_flat_scaling.py
n= 1,000 0.85 ms/query
n= 5,000 4.79 ms/query
n= 10,000 8.50 ms/query
n= 50,000 45.79 ms/query
n=100,000 98.35 ms/query
n=500,000 658.69 ms/query
n=1,000,000 1258.07 ms/query
To do better, we need to prune the search to only vectors likely to be close to the query.
Aside: Curse of dimensionality
In low dimensions, a natural way to prune the search space is spatial partitioning. A QuadTree recursively divides 2D space into cells. To find the nearest neighbor of a query, locate its cell and only check the vectors inside. An excellent visualization is available here.
+--------------------+------------------------+
| | |
| · · · | · · |
| | · | (skip)
| · | · |
| | |
+----------+---------+------------------------+
| · · | · [Q] | |
| · | · · | · · | (search Q's cell only)
| | · | |
+----------+---------+------------------------+
| · · | · · |
| · | · | (skip)
+--------------------+------------------------+
[Q] = query · = indexed vectorThis doesn’t generalise to high dimensions. The natural extension splits every cell along all axes at once, producing sub-regions per split. But is also exactly the number of distinct binary vectors in dimensions, so after a single split, each cell holds at most one data point and the index offers no pruning at all. For continuous embeddings the same logic applies: at you already have a billion cells, more than most datasets have vectors, and the index degenerates into a linear scan.
A smarter approach, the Kd-tree, splits along one dimension at a time and stays shallow. But even this fails at high : the deeper problem is that high-dimensional space is mostly empty. As grows, all points drift to roughly the same distance from any query, and there is no “nearby” region worth exploiting. This is the curse of dimensionality.
Scaling with IVF (Inverted File Index)
IVF sidesteps the curse of dimensionality by learning the partitions from data. K-means clustering finds centroids that summarise the natural groupings in the corpus. Unlike axis-aligned splits, these clusters naturally group vectors in high dimensions.
Building the index is two steps:
- Train: run k-means on the corpus to find
nlistcentroids. - Index: assign every vector to its nearest centroid.
Each centroid defines a Voronoi cell. This is the region of space closer to it than to any other centroid.

At query time, find the nprobe nearest centroids and run exact search only within those clusters. Higher nprobe improves recall but slows down search, taking us to the core accuracy/speed tradeoff in IVF. OpenSearch for example exposes both nlist and nprobe as first-class IVF parameters.
class IVFIndex:
def __init__(self, nlist: int, nprobe: int):
self._nlist = nlist
self._nprobe = nprobe
self._centroids = None
self._lists = {} # cluster_id → [(vector, id)]
def train(self, vectors: np.ndarray):
# Random initialisation
centroids = np.random.default_rng().choice(vectors, size=self._nlist, replace=False, axis=0)
for _ in range(25):
# Assign each vector to its nearest centroid
labels = np.argmin(np.linalg.norm(vectors[:, None] - centroids[None], axis=2), axis=1)
# Recompute centroids as the mean of their assigned vectors
new_centroids = np.array([
vectors[labels == i].mean(axis=0) if (labels == i).any() else centroids[i]
for i in range(self._nlist)
])
# End loop early if the centroids converge
if np.allclose(centroids, new_centroids):
break
centroids = new_centroids
self._centroids = centroids
def index(self, vector_id: int, vector: np.ndarray):
cell = int(np.argmin(np.linalg.norm(self._centroids - vector, axis=1)))
self._lists.setdefault(cell, []).append((vector, vector_id))
def search(self, query: np.ndarray, k: int) -> list[tuple[int, float]]:
# 1. Find the nprobe nearest centroids
cells = np.argpartition(np.linalg.norm(self._centroids - query, axis=1), self._nprobe)[:self._nprobe]
# 2. Gather candidates from those clusters
candidates, ids = [], []
for cell in cells:
for vec, vid in self._lists.get(int(cell), []):
candidates.append(vec)
ids.append(vid)
# 3. Exact search within candidates only
distances = np.linalg.norm(np.stack(candidates) - query, axis=1)
nearest = np.argpartition(distances, k)[:k]
nearest = nearest[np.argsort(distances[nearest])]
return [(ids[i], float(distances[i])) for i in nearest]IVF in action: a worked example
To make this concrete, consider 8 vectors in 2D and a query , with nlist=3 and nprobe=1. We obtain 3 clusters:
Vectors: v0=[1,2] v1=[2,1] v2=[4,3] (cluster A)
v3=[8,9] v4=[9,8] v5=[8.5,8.5] (cluster B)
v6=[5,1] v7=[6,2] (cluster C)
After k-means training:
Centroid A = [2.3, 2.0]
Centroid B = [8.5, 8.5]
Centroid C = [5.5, 1.5]Note that is geometrically close to cluster C, but k-means assigned it to A (dist to A=1.97, dist to C=2.12). This is a borderline point.
At query time, we compute the distance from to each centroid:
dist(q, A) = √((5-2.3)² + (4-2.0)²) = 3.36
dist(q, B) = √((5-8.5)² + (4-8.5)²) = 5.70
dist(q, C) = √((5-5.5)² + (4-1.5)²) = 2.55 ← nearestWith nprobe=1, we only search cluster C: vectors (dist=3.0) and (dist=2.24). IVF returns as the nearest neighbor. But the true nearest neighbor is at dist==1.41, sitting in cluster A. We never look there.
Setting nprobe=2 would search clusters C and A, finding . This is the fundamental IVF trade-off: nprobe=1 checked 2 out of 8 vectors (75% pruned) but missed the best match. nprobe=nlist degenerates to a flat scan, which is correct but slow.

Tuning IVF parameters
-
nlist(number of clusters): Controls the granularity of partitioning. Too few clusters and each one is large, so you don’t prune much. Too many and clusters become sparse, with nearby vectors split across boundaries. A common starting point is for vectors: 1,000 clusters for a million vectors. -
nprobe(clusters to search): The accuracy/speed dial. Start with (5% of clusters) and increase until recall plateaus. In practice, searching 5-10% of clusters often reaches 95%+ recall.
Benchmarking IVF against the flat index on 128-dimensional clustered data (using nlist and nprobe of nlist) shows the latency benefits:
> uv run plot_ivf_performance.py
n= 1,000 | Flat= 1.12 ms | IVF= 0.24 ms
n= 5,000 | Flat= 5.77 ms | IVF= 2.11 ms
n= 10,000 | Flat= 11.20 ms | IVF= 1.70 ms
n= 25,000 | Flat= 29.05 ms | IVF= 4.27 ms
n= 50,000 | Flat= 66.09 ms | IVF= 5.36 ms
n=100,000 | Flat= 157.11 ms | IVF= 10.53 ms
At 100k vectors, IVF is 15x faster. The gap widens with scale: the time taken for flat index grows linearly while IVF only scans a fixed fraction of the data. But speed alone does not matter if you are returning the wrong results. Sweeping nprobe on the 100k index reveals the trade-off:
nprobe= 1 ( 0.3%) | Recall=0.588 | 2.37 ms
nprobe= 4 ( 1.3%) | Recall=0.986 | 3.19 ms
nprobe= 16 ( 5.1%) | Recall=1.000 | 12.27 ms
nprobe=316 (100.0%) | Recall=1.000 | 237.25 ms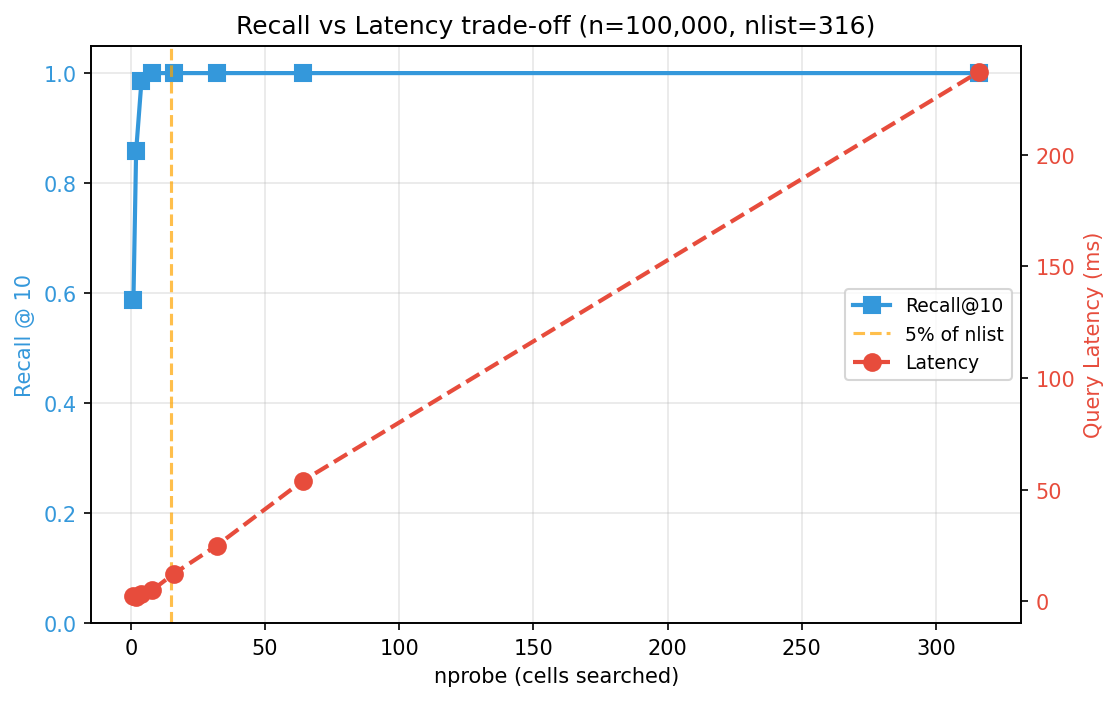
Recall climbs steeply at first (probing just 1.3% of clusters already hits 98.6% recall) and then flatlines while latency keeps growing linearly. Past the knee of the curve, every additional cluster probed costs time without improving results. Finding that optimal value of nprobe is key.
HNSW
Hierarchical Navigable Small World (HNSW) requires no pre-training and no centroid count to tune. The implementation follows the original paper.

The name describes the structure:
- Small World: every node connects to its
Mnearest neighbors, forming a graph where any two nodes are reachable in hops, a property of navigable small-world networks. HigherMmeans a better-connected graph (better recall, more memory). - Navigable: search is a greedy walk: at each hop, move to the neighbor closest to the query. The
efparameter controls how many candidates to track during this walk:ef=1is pure greedy, while higherefexplores more paths for better accuracy. - Hierarchy: multiple layers stacked on the small world graph: sparse at the top for fast coarse navigation, and dense at the bottom for precision. Each node appears at a random layer drawn from an exponential distribution; most nodes only exist at layer 0, a few reach higher layers.
If you’ve seen a skip list, this is the same idea generalised to vectors: express lanes at the top let you take long hops toward the target, then you drop to denser layers for precision. The difference is that a skip list’s “closeness” is a 1D key comparison; HNSW’s is a vector distance.
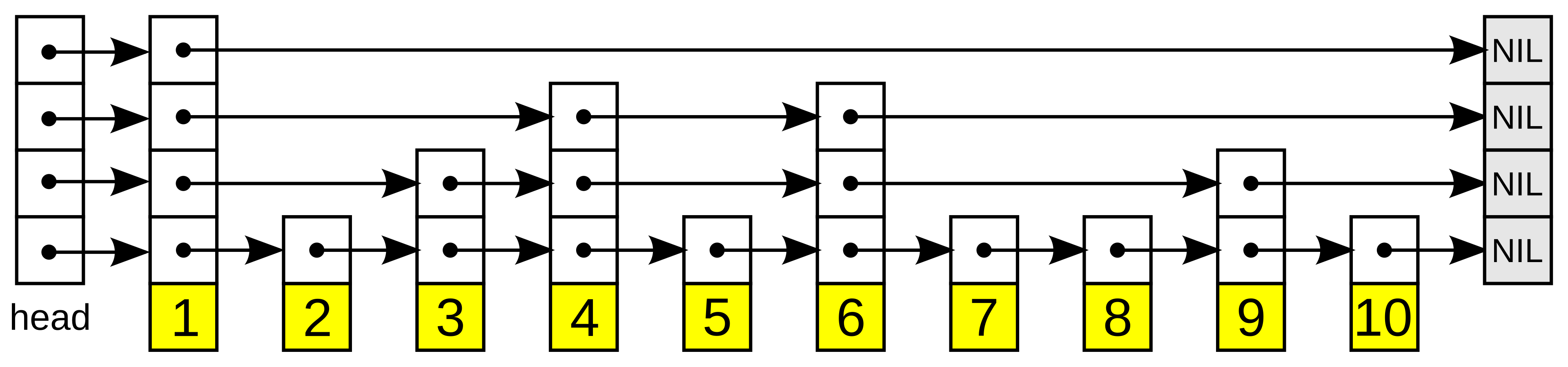
Search enters at the top layer, greedily descends toward the query, then repeats on each denser layer until layer 0. The core primitive is a greedy best-first search with a bounded candidate set: expand the closest unvisited node, but skip any neighbor that cannot improve on the current ef best results. With ef=1 this is pure greedy search; with ef>1 it becomes a beam search that tracks multiple promising paths. This bounds exploration to a local neighborhood rather than the full graph.
def _search_layer(self, query: np.ndarray, entry_ids: list[int], ef: int, layer: int) -> list:
# Seed the candidate min-heap with all entry points
candidates = [] # min-heap: (dist, id): closest node explored next
for node_id in entry_ids:
dist = np.linalg.norm(query - self._nodes[node_id].vector)
heapq.heappush(candidates, (dist, node_id))
best = [] # max-heap (Python 3.14+): worst of current top-ef results at front
visited = set()
while candidates:
d, node_id = heapq.heappop(candidates)
if node_id in visited:
continue
visited.add(node_id)
heapq.heappush_max(best, (d, node_id))
if len(best) > ef:
heapq.heappop_max(best) # evict the furthest if over budget
for nb_id in self._nodes[node_id].neighbors.get(layer, []):
nb_dist = np.linalg.norm(query - self._nodes[nb_id].vector)
if len(best) < ef or nb_dist < best[0][0]: # only explore if promising
heapq.heappush(candidates, (nb_dist, nb_id))
return sorted(best) # closest firstThe index itself holds two things: the nodes (each storing a vector and a per-layer neighbor list), and bookkeeping for the current entry point and top layer. M, ef_construction, and ef_search are the HNSW parameters OpenSearch exposes for tuning.
@dataclass
class HNSWNode:
vector: np.ndarray
neighbors: dict[int, list[int]] = field(default_factory=dict) # layer → [node_ids]
class HNSWIndex:
def __init__(self, M: int = 16, ef_construction: int = 200, ef_search: int = 50):
self._M = M # max neighbors per node per layer
self._ef_construction = ef_construction # beam width during insert
self._ef_search = ef_search # beam width during search
self._nodes: dict[int, HNSWNode] = {}
self._entry_point: int | None = None
self._max_layer: int = 0
self._mL = 1 / np.log(M) # layer probability scale
def _random_level(self) -> int:
# Exponential distribution: most nodes at layer 0, exponentially fewer above
return int(-np.log(np.random.uniform()) * self._mL)
def index(self, vector: np.ndarray) -> None:
node_id = len(self._nodes)
node = HNSWNode(vector=vector)
self._nodes[node_id] = node
node_layer = self._random_level()
if self._entry_point is None:
self._entry_point = node_id
self._max_layer = node_layer
return
# Phase 1: ef=1 greedy descent through layers above node_layer: find a close entry point
entry_ids = [self._entry_point]
for layer in range(self._max_layer, node_layer, -1):
entry_ids = [nid for _, nid in self._search_layer(vector, entry_ids, ef=1, layer=layer)]
# Phase 2: beam search at each layer from node_layer down to 0: wire edges
for layer in range(min(node_layer, self._max_layer), -1, -1):
M_limit = self._M * 2 if layer == 0 else self._M
results = self._search_layer(vector, entry_ids, ef=self._ef_construction, layer=layer)
entry_ids = [nid for _, nid in results]
# Connect new node to its M nearest neighbors at this layer
node.neighbors[layer] = [nid for _, nid in heapq.nsmallest(M_limit, results)]
# Bidirectional: each neighbor adds a back-edge, then prunes to M
for nb_id in node.neighbors[layer]:
nb = self._nodes[nb_id]
all_edges = nb.neighbors.get(layer, []) + [node_id]
scored = [(np.linalg.norm(nb.vector - self._nodes[n].vector), n) for n in all_edges]
nb.neighbors[layer] = [nid for _, nid in heapq.nsmallest(M_limit, scored)]
if node_layer > self._max_layer:
self._max_layer = node_layer
self._entry_point = node_id
def search(self, query: np.ndarray, k: int) -> list[tuple[int, float]]:
# ef=1 descent to layer 1: fast coarse navigation
entry_ids = [self._entry_point]
for layer in range(self._max_layer, 0, -1):
entry_ids = [nid for _, nid in self._search_layer(query, entry_ids, ef=1, layer=layer)]
# Full beam search at layer 0: precision
return self._search_layer(query, entry_ids, ef=self._ef_search, layer=0)[:k]HNSW in action: a worked example
Same 8 vectors as before: , , , , , , , . Suppose the graph has been built with M=2:
Layer 1 (sparse): v3 ——— v0
Layer 0 (dense): v0 — v1 — v2
| |
v6 — v7 v3
\ |
v4—v5To search for with k=1:
Phase 1: Layer 1 (ef=1): Start at entry point (dist=5.83). Its only layer-1 neighbor is (dist=4.47). is closer, so it becomes the entry. Descend to layer 0.
Phase 2: Layer 0 (ef=ef_search): Start at (dist=4.47). Expand its neighbors: (4.24), (3.0). Pop closest candidate , expand: (2.24). Pop , expand: (5.66, no improvement). Pop , expand: (1.41). is now the closest candidate. Expand : (5.83, no improvement). No candidates left that can beat 1.41. Return .
HNSW found the exact nearest neighbor by visiting most of the 8 nodes. In a toy graph that is expected, but the structure scales logarithmically: at a million vectors with M=16, a search typically visits a few hundred nodes.
Tuning HNSW parameters
-
M(edges per node): Controls graph connectivity. HigherMmeans more edges, better recall, but more memory. Each doubling ofMroughly doubles the index size.M=16is a solid default; increase to 32-64 for datasets where recall matters more than memory. -
ef_construction(beam width during index building): How many candidates to consider when connecting a new node. Higher values produce a better-connected graph at the cost of slower indexing.ef_construction=200is a common default; for large production indexes, 100-500 is typical. This only affects build time, not search. -
ef_search(beam width during query): The runtime accuracy/speed knob, analogous to IVF’snprobe. Must be . Start at 50 and increase until recall stabilizes. UnlikeMandef_construction, this can be tuned per-query without rebuilding the index.
Benchmarking HNSW
Our pure-Python implementation above is faithful to the paper, but far too slow for realistic benchmarks. Indexing 100k vectors takes minutes instead of seconds. The numbers below use hnswlib, a production-grade C++ implementation of the same algorithm, with the same M, ef_construction, and ef_search parameters. On the same clustered dataset:
> uv run plot_hnsw_performance.py
n= 1,000 | Flat= 1.54 ms | IVF= 0.51 ms | HNSW= 0.058 ms
n= 5,000 | Flat= 5.36 ms | IVF= 0.96 ms | HNSW= 0.042 ms
n= 10,000 | Flat= 12.10 ms | IVF= 1.57 ms | HNSW= 0.046 ms
n= 25,000 | Flat= 28.73 ms | IVF= 2.46 ms | HNSW= 0.060 ms
n= 50,000 | Flat= 64.24 ms | IVF= 6.55 ms | HNSW= 0.073 ms
n=100,000 | Flat= 143.35 ms | IVF= 11.15 ms | HNSW= 0.095 ms
At 100k vectors, HNSW queries take 0.095ms, which is 1500x faster than flat search and 100x faster than IVF. The scaling is visible: latency barely moves as the corpus grows from 1k to 100k.
Sweeping ef_search on the 100k index shows the same recall/latency pattern as IVF’s nprobe:

At ef_search=50, recall is 96.8% at 0.115ms per query. Doubling to ef_search=100 reaches 99.6% at 0.160ms. The full useful range fits within sub-millisecond latency.
The recall trade-off
Every approximate algorithm sacrifices some accuracy for speed. Recall@k measures this: of the true nearest neighbors, what fraction does the algorithm actually return?
A recall of 0.95 means the algorithm finds 95 of the true 100 nearest neighbors. The remaining 5 are close. They’re near-misses, not random vectors from the other side of the space. For most applications (RAG, recommendations, semantic search), this is indistinguishable from exact search.
The relationship between recall and speed is different for each algorithm:
| Flat | IVF | HNSW | |
|---|---|---|---|
| Recall | 1.0 (exact) | Controlled by nprobe | Controlled by ef_search |
| Search time | |||
| Index memory | + centroids | ||
| Build time | None | K-means training | Incremental (no batch step) |
| Sweet spot | Large , memory-constrained | When recall and speed both matter |
In practice, HNSW typically achieves 95%+ recall while searching less than 1% of the dataset. IVF can match this recall but needs a well-chosen nlist and nprobe, a training step that HNSW avoids entirely. The flat index remains the right choice for small datasets or when you need guaranteed exact results.
Conclusion
These algorithms are building blocks, not final answers. Production systems layer them: IVF with product quantization for compression, HNSW with metadata pre-filtering, or hybrid approaches that use IVF to coarsely shard the graph. The ANN Benchmarks project tracks how these combinations perform across real datasets, and it is the best place to start when choosing an index for a specific workload.